Access a toolkit of in vitro models to study mechanisms underlying motor neuron diseases
Powered by opti-ox
Powered by opti-ox
Lower motor neurons are a diverse group of neuronal subtypes that regulate muscle and glandular activity. They form complex neural circuits by integrating inputs from interneurons, sensory neurons, and upper motor neurons to control behaviours such as locomotion. Disruption of these networks leads to motor neuron diseases (MNDs), including spinal muscular atrophy (SMA) and amyotrophic lateral sclerosis (ALS).
The underlying causes of MNDs are still not fully understood, so replicating the slow degeneration of motor neurons in vitro remains a challenge. Moreover, preclinical animal models often fail to capture human disease biology, while differentiating and culturing human iPSC-derived motor neurons is a complex process, and published protocols show cells clumping when cultured1-4. This excessive clumping impacts the quality of downstream assays. ioMotor Neurons provide scientists with a reliable, functional source of defined lower motor neurons, ensuring clump-free cultures and single-cell resolution.
Traditional human iPSC-derived motor neurons that clump and aggregate make high-content imaging challenging. When cells pile up, signals are weaker and more cells are required per well.
ioMotor Neurons rapidly acquire a homogeneous motor neuron phenotype upon thawing and maintain a single cell distribution during maturation, even after 41 DIV, giving scientists confidence in reliable long-term cultures.
The absence of cell clusters improves MEA, whole cell patch clamp and axon tracking assays by enabling:
In this video, Dr Ben Bar-Sadeh, Anima Biotech focuses on his team's efforts to use an AI-powered, visual biology approach for drug discovery. When imaging healthy ioMotor Neurons alongside their genetically-matched TDP-43 and SOD1 disease models, immunofluorescence reveals distinct phenotypic differences. A proprietary marker signal is significantly elevated with visible aggregates in the TDP-43 model and markedly decreased in the SOD1 model, providing insights into disease-specific cellular changes.

High-resolution confocal imaging validates the protocol for co-culturing ioSkeletal Myocytes with ioMotor Neurons. By day 30, the system displays distinct structural organisation, with MAP2-positive neurons (red) in the culture alongside Desmin-positive myocytes (cyan). The presence of acetylcholine receptors (yellow, alpha-bungarotoxin) confirms that this co-culture method supports the development of key neuromuscular features suitable for complex assay development.
In this video, our scientist takes you through the step-by-step process of how to thaw, seed and culture ioMotor Neurons.

Dr Irit Reichenstein
Senior Scientist | Anima Biotech

Dr Elizabeth Di Lullo
Associate Scientific Director | Brainever


Access 14 ALS and FTD disease models with disease-related mutations such as SOD1, FUS, MAPT and TDP-43 (TARDBP) genetically engineered in ioGlutamatergic Neurons and ioMotor Neurons.


Interested in gene knockouts and CRISPR screens?
CRISPRko-Ready ioMotor Neurons cells engineered to constitutively express Cas9 nuclease for the quick and easy generation of gene knockouts and CRISPR screens.

Build your custom disease model or reporter line to pair with wild-type ioMotor Neurons as the genetically matched control.
Throughout the custom process, our experts will bring your project to life, and be on hand to support you with any technical queries.
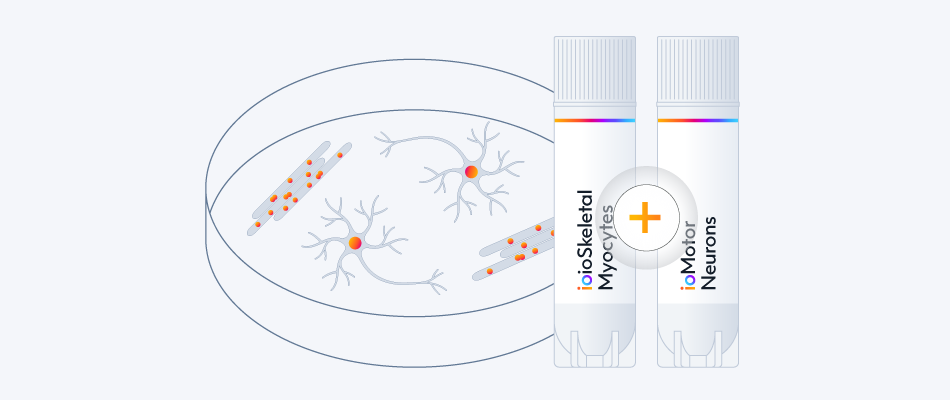

Study neuromuscular interactions and the impact of ALS-disease-related mutations by co-culturing ioMotor Neurons with skeletal myocytes. Access 14 disease models and the single co-culture protocol.
View the co-culture protocol
Explore ALS & FTD Disease Models
Explore ioSkeletal Myocytes
bit.bio
bit.bio
Vaquero, et al
bit.bio
2023
Foulser, et al
bit.bio
2024
Brown et al.
bit.bio
2024
Smith et al.
bit.bio
2024
Veteleanu et al.
bit.bio
2025
Foulser et al.
bit.bio
2025
Hill et al.
bit.bio
2026
Davenport A, Frolov T & Kotter M
Drug Discovery World
2020
Filippa VG, et al.
bioRxiv
2026
Using ioGlutamatergic Neurons and ioMotor Neurons
Dr Elizabeth Di Lullo | Associate Scientific Director | BrainEver
Human Cell Forum 2025
Session 1 Track 1 | Modelling neurodegeneration in vitro with human iPSC-derived cells
Dr Grace Cooper | Senior Scientist | bit.bio
Human Cell Forum 2025
Session 1 Track 2 | From cells to systems: Building human iPSC-derived models of pain, neuromuscular junctions, and glial dynamics
Dr Ben Bar-Sadeh | Senior Scientist | Anima Biotech
Human Cell Forum 2025
Session 3 | Making complex human biology compatible with modern drug discovery workflows
bit.bio
Luke Foulser | Scientist | bit.bio
Prof Roger Pedersen | Adjunct Professor and Senior Research Scientist at Stanford University
Dr Thomas Moreau | Director of Cell Biology Research | bit.bio
Tom Brown | Senior Product Manager | bit.bio
Marcos Herrera Vaquero, PhD | Senior Scientist | bit.bioBen Bar-Sadeh, PhD | Senior Scientist | Anima Biotech
Tom Brown | Senior Product Manager | bit.bio
Lower motor neurons are a diverse class of cells that serve as the essential regulators of skeletal muscle activity and glandular activity. By integrating complex inputs from sensory neurons, interneurons, and upper motor neurons, they orchestrate voluntary behaviours such as movement. Consequently, motor neuron degeneration drives debilitating Motor Neuron Diseases (MNDs) such as Spinal Muscular Atrophy (SMA) and Amyotrophic Lateral Sclerosis (ALS).
ioMotor Neurons address the technical challenge of cell clumping often seen in traditional differentiation protocols by maintaining a homogenous, single-cell monolayer distribution. This clump-free monolayer improves quality on downstream assays. Powered by opti-ox, these human iPSC-derived motor neurons provide scientists a consistent source of cells ideal for disease modelling.
ioMotor Neurons improve microelectrode array (MEA) data quality by maintaining a clump-free, single-cell monolayer that ensures direct contact with the recording electrodes. This precise coupling provides a better signal-to-noise ratio, allowing for the clear isolation of single neurons and enhancing the accuracy of functional network analysis.
ioMotor Neurons engineered with ALS-relevant mutations, such as those in the SOD1 or TARDBP (TDP-43) genes, exhibit distinct disease-specific phenotypes compared to genetically matched controls. Validated applications include quantifying elevated protein aggregates (in TARDBP models) or marker downregulation (in SOD1 models), providing a robust, human-relevant platform for phenotypic screening and drug discovery.
Co-culturing ioMotor Neurons with ioSkeletal Myocytes enables the in vitro reconstruction of functional neuromuscular units. Confocal imaging confirms the development of structural organisation, including the presence of acetylcholine receptors (alpha-bungarotoxin positive), allowing scientists to investigate the specific crosstalk between motor neurons and muscle cells implicated in neuromuscular diseases.