






cat no | io1055
ioMotor Neurons FUS P525L/WT
Human iPSC-derived ALS disease model
-
Cryopreserved human iPSC-derived cells powered by opti-ox, that are ready for experiments in days
-
Engineered to enable investigations into the impact of mutant FUS protein on neurodegenerative disease
-
Clump free, highly-pure motor neurons, that form functional neuronal networks in co-culture with astrocytes

Human iPSC-derived ALS disease model

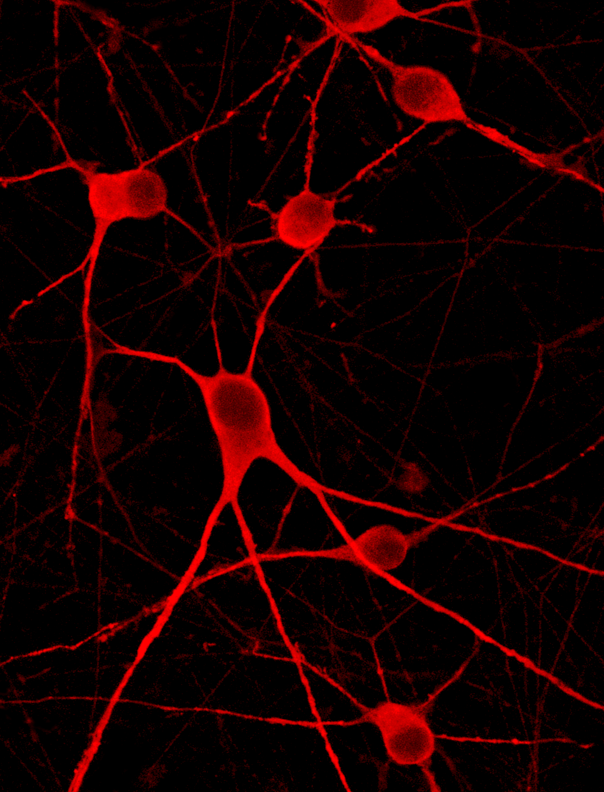
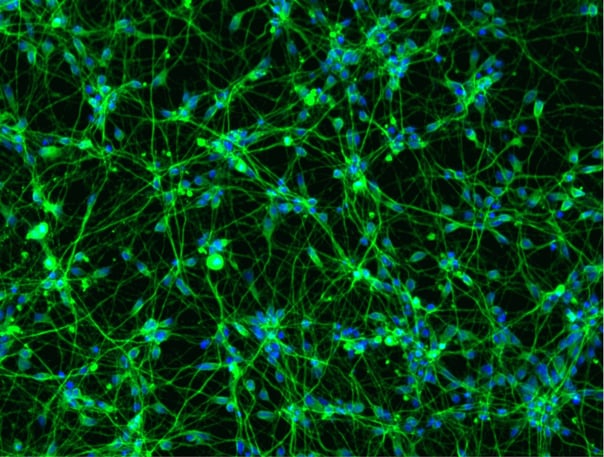
ioMotor Neurons FUS P525L/WT form a homogenous neuronal network by day 4
ioMotor Neurons FUS P525L/WT rapidly acquire a motor neuronal phenotype, forming homogenous neuronal networks, without clumping of cells. Compared to the genetically matched wild type control, ioMotor Neurons. Day 1 to 11 post thawing; 100X magnification.

ioMotor Neurons FUS P525L/WT express motor neuron-specific markers with protein expression highly reminiscent to the genetically matched control
Immunofluorescent staining on post-revival day 11 demonstrates similar homogenous expression of pan-neuronal protein TUBB3, motor neuron specific marker ISL2 and the cholinergic marker VAChT in ioMotor Neurons FUS P525L/WT compared to the genetically matched control, ioMotor Neurons.

ioMotor Neurons FUS P525L/WT express motor neuron-specific markers with protein expression highly reminiscent to the genetically matched control
Immunofluorescent staining on post-revival day 11 demonstrates similar homogenous expression of pan-neuronal protein MAP2, motor neuron specific marker HB9 and the cholinergic marker VAChT in ioMotor Neurons FUS P525L/WT compared to the genetically matched control, ioMotor Neurons.

Disease-related FUS is expressed in ioMotor Neurons FUS P525L/WT following deterministic programming
Gene expression analysis demonstrates that ioMotor Neurons FUS P525L/WT and the genetically matched control (WT) express the FUS gene encoding the Fused in Sarcoma protein. Gene expression levels were assessed by RT-qPCR (data expressed relative to the parental hiPSC control (iPSC Control), normalised to HMBS). Data represents day 11 post-revival samples.

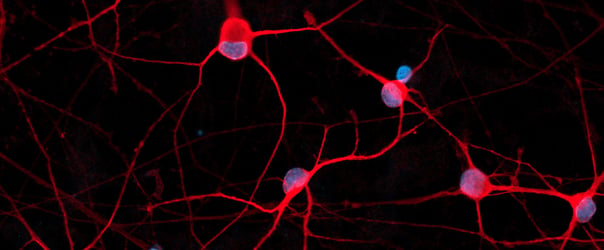
Co-culture of ioMotor Neurons and ioSkeletal Myocytes
High resolution confocal imaging of ioSkeletal Myocytes (io1002) and ioMotor Neurons wild-type co-culture. Staining with alpha-bungarotoxin (yellow) highlights acetylcholine receptor expression on co-cultured ioSkeletal Myocytes. Desmin (cyan) and microtubule-associated protein 2 (red) define ioSkeletal Myocytes and ioMotor Neurons respectively. Co-culture imaged at day 30, 40X magnification.
Download the step-by-step protocol for culturing ioSkeletal Myocytes and ioMotor Neurons.

Do more with every vial
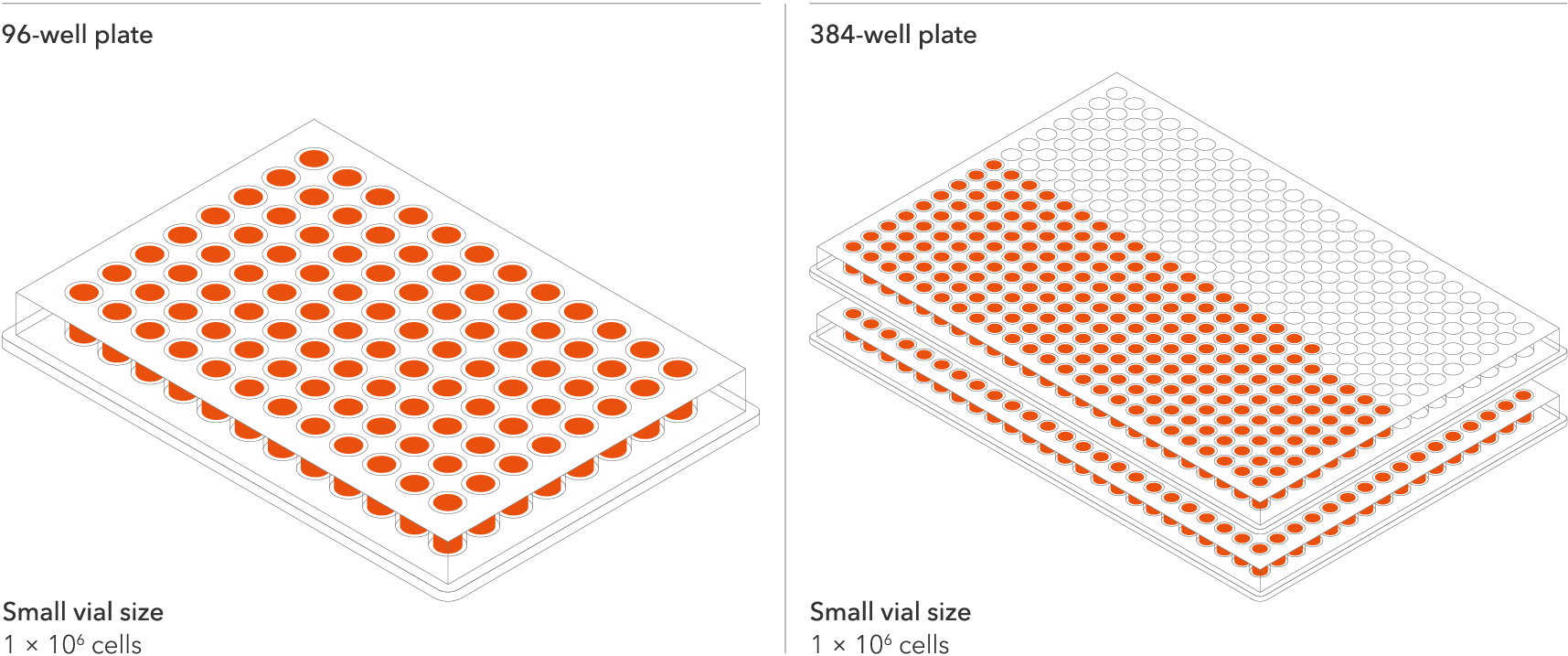
The seeding density of our human iPSC-derived ioMotor Neurons and related disease models has been optimised and validated to a recommended seeding density of 30,000 cells/cm². This means scientists can do more with every vial and expand experimental design within budget without losing out on quality. Resulting in more experimental conditions, more repeats, and more confidence in the data. One Small vial can plate a minimum of 0.7 x 24-well plate, 1 x 96-well plate, or 1.5 x 384-well plate.
Vial limit exceeded
A maximum number of 20 vials applies. If you would like to order more than 20 vials, please contact us at orders@bit.bio.








Hoescht(blue)TUBB3(blue)_day4.png?width=604&name=bit.bio_ioGlutamatergic%20Neurons_60xMAP2(red)Hoescht(blue)TUBB3(blue)_day4.png)








.png?width=604&name=ioMotor%20Neuron%20Hero%20image%20for%20Webinar%20(1).png)