




















cat no | io1007
ioGlutamatergic Neurons GBA null/R159W
Human iPSC-derived Gaucher and Parkinson’s disease model
-
Cryopreserved human iPSC-derived cells powered by opti-ox that are ready for experiments in days
-
Engineered to carry a GBA mutation relevant for modelling Gaucher and Parkinson's diseases
-
Consistent, functional excitatory neurons that form neuronal networks within days

Human iPSC-derived Gaucher and Parkinson’s disease model

Reduced glucocerebrosidase activity demonstrated in disease model cells carrying GBA mutations
Glucocerebrosidase (GCase) activity was measured in ioGlutamatergic Neurons carrying mutations in the GBA gene (GBA null/R159W, GBA null/null and GBA null/N409S) or carrying SNCA A53T mutations (SNCA A53T/A53T and SNCA A53T/WT), and the wild-type genetically matched control.
Cells were seeded in 96-well plates (25,000 cells per well) and a fluorometric assay using 4-MUG as substrate was carried out on DIV9 to measure the enzymatic activity of GCase.
The assay confirmed that iPSC-derived glutamatergic neurons with GBA mutations showed significantly reduced GCase activity compared to the wild-type control and SNCA A53T disease models.
Statistical analysis: One-way ANOVA followed by Dunnett’s multiple comparison test versus isogenic control ***p<0.001.
Data courtesy of T. Loeffler et al., Scantox

GBA protein is present in ioGlutamatergic Neurons GBA null/R159W at a lower level than the wild type control
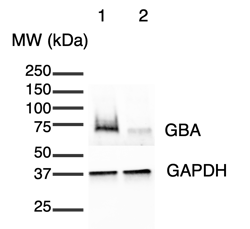
A Western blot experiment confirmed the presence of the GBA protein in ioGlutamatergic Neurons GBA null/R159W at a lower level than in the wild-type ioGlutamatergic Neurons. Day 11 cell lysates were subjected to Western blotting (20 µg protein in 40 µl per lane) using 4-20% mini protean TGX stain-free gels. Proteins were transferred onto PVDF membranes using the Trans-Blot Turbo Transfer Pack, blocked for 10 minutes, incubated with primary antibodies (GBA Invitrogen MA5-26589, 1:2000; GAPDH Abcam ab8245, 1:5000), washed three times, incubated with HRP-labelled secondary antibodies, washed three times and signal visualised by electrochemiluminescence.
1= ioGlutamatergic Neurons (wild type), 2= ioGlutamatergic Neurons GBA null/R159W.

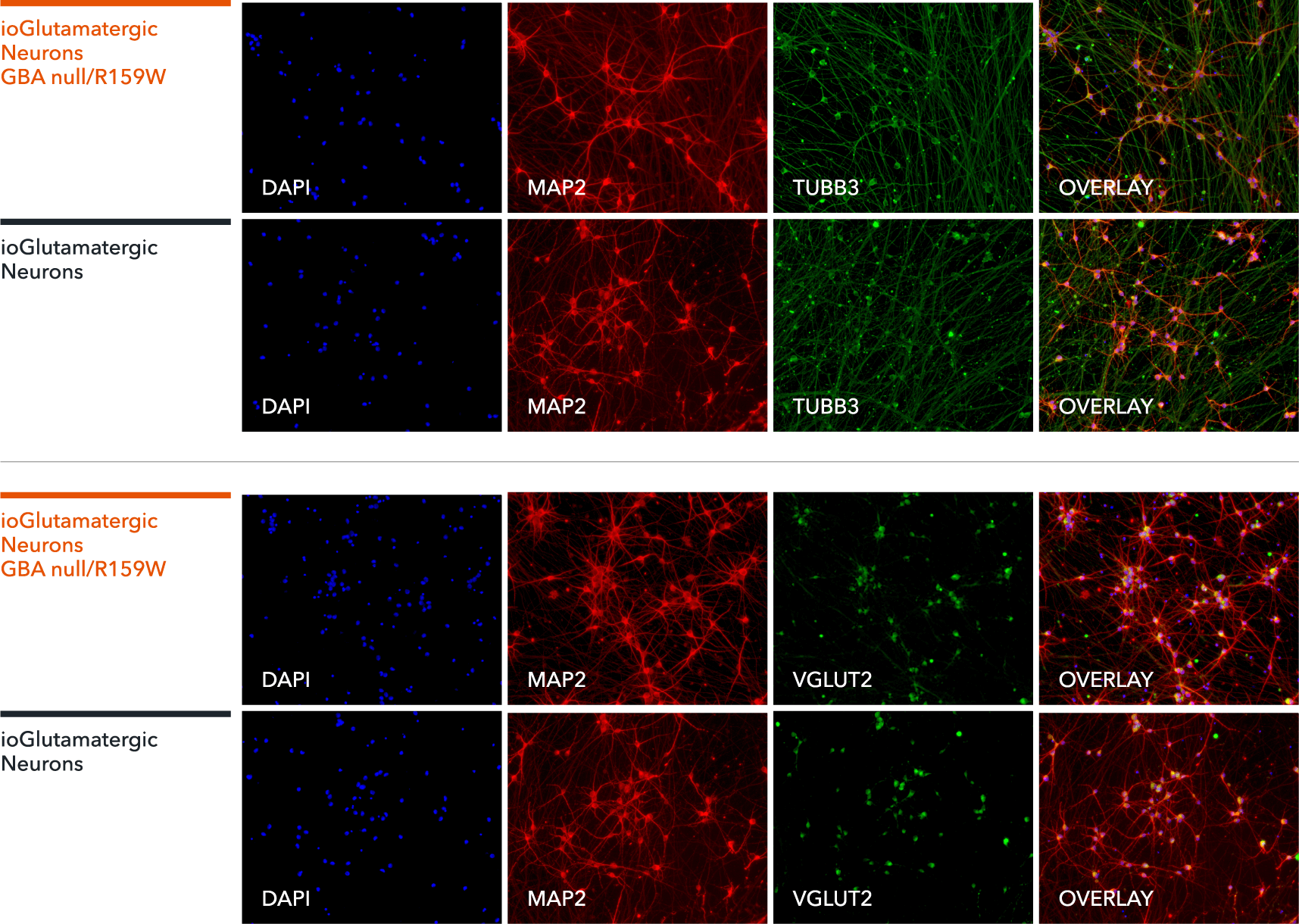
ioGlutamatergic Neurons GBA null/R159W express neuron-specific markers comparably to the isogenic control
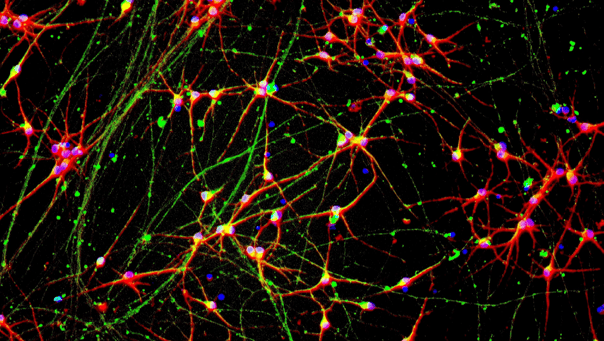
Immunofluorescent staining on post-revival day 11 demonstrates similar homogenous expression of pan-neuronal proteins TUBB3 and MAP2 (upper panel) and glutamatergic neuron-specific transporter VGLUT2 (lower panel) in ioGlutamatergic Neurons GBA null/R159W compared to the isogenic control. 100X magnification.

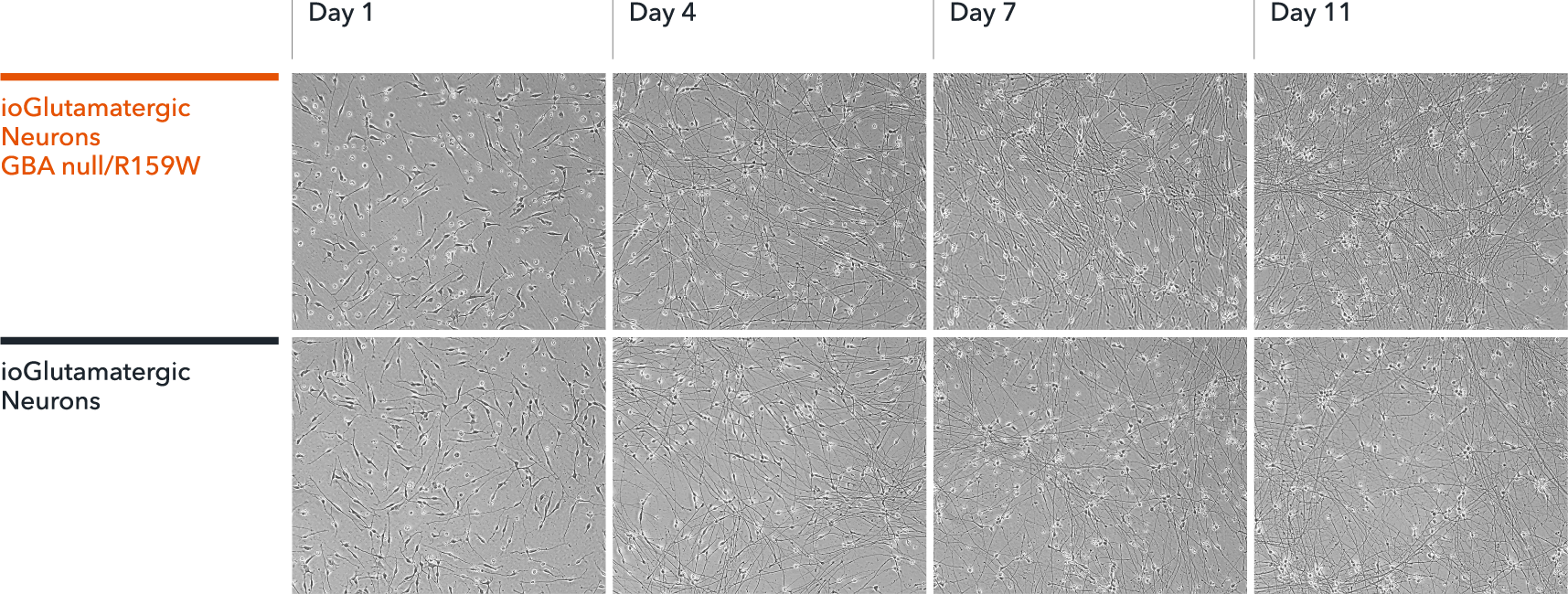
ioGlutamatergic Neurons GBA null/R159W form structural neuronal networks by day 11
ioGlutamatergic Neurons GBA null/R159W mature rapidly and form structural neuronal networks over 11 days, when compared to the isogenic control. Day 1 to 11 post thawing; 100X magnification.

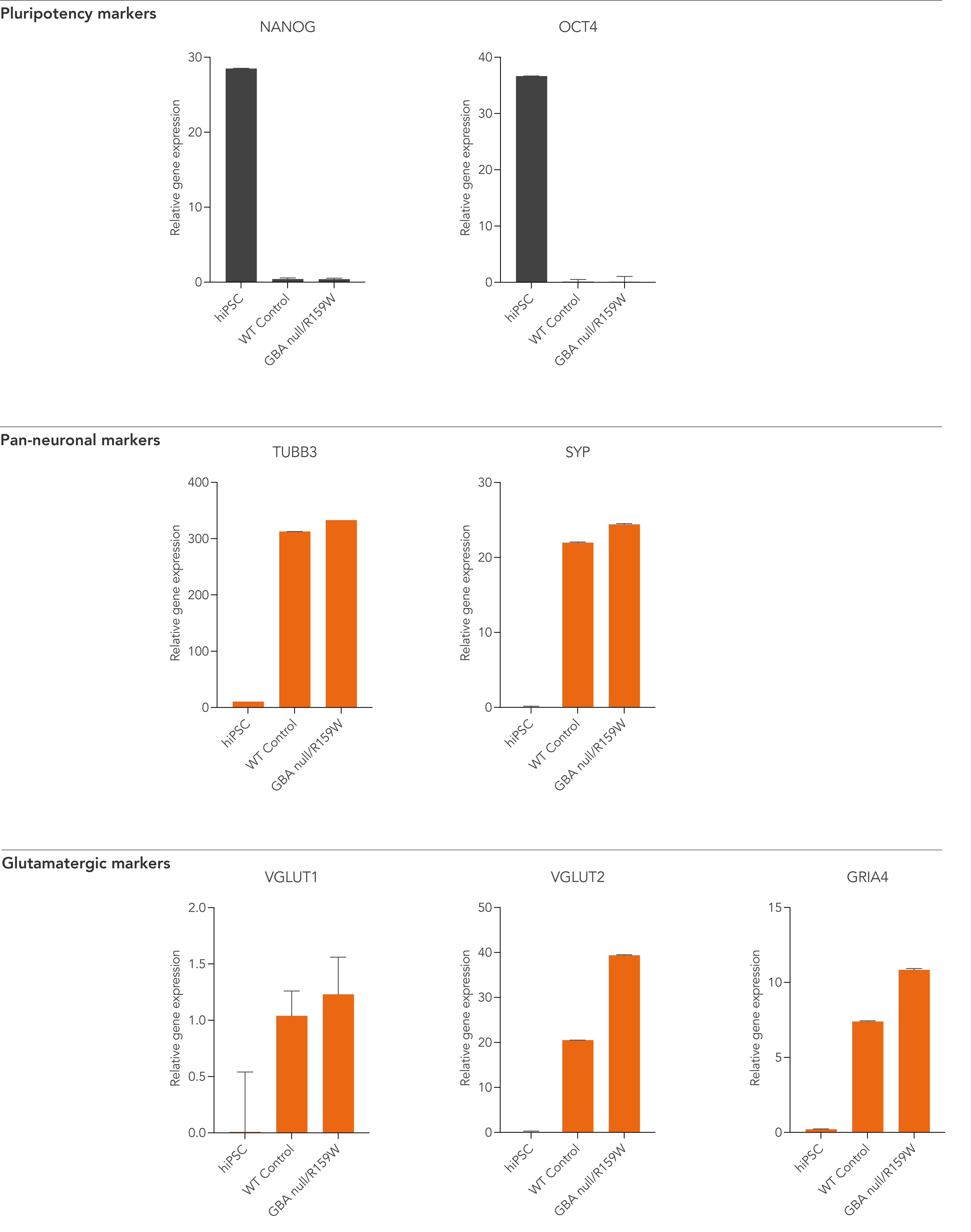
ioGlutamatergic Neurons GBA null/R159W demonstrate gene expression of neuronal and glutamatergic-specific markers following deterministic programming
Gene expression analysis demonstrates that ioGlutamatergic Neurons GBA null/R159W and the isogenic control (WT Control) lack the expression of pluripotency markers (NANOG and OCT4) at day 11, whilst robustly expressing pan-neuronal (TUBB3 and SYP) and glutamatergic specific (VGLUT1 and VGLUT2) markers, as well as the glutamate receptor GRIA4. Gene expression levels were assessed by RT-qPCR (data normalised to HMBS; cDNA samples of the parental human iPSC line (hiPSC) were included as reference). Data represents day 11 post-revival samples, n=2 replicates.
View the step-by-step RNA extraction and RT-qPCR protocol used to generate this data

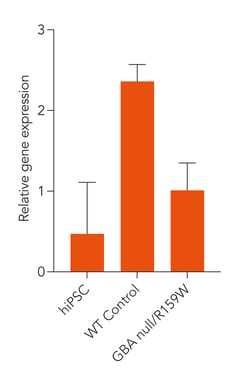
Disease-related GBA is expressed in ioGlutamatergic Neurons GBA null/R159W following deterministic programming
Gene expression analysis demonstrates that ioGlutamatergic Neurons GBA null/R159W and the isogenic control (WT Control) express the GBA gene encoding the glucocerebrosidase protein. Gene expression levels were assessed by RT-qPCR (data normalised to HMBS, cDNA samples of the parental human iPSC line (hiPSC) were included as reference). Data represents day 11 post-revival samples, n=2 replicates.

Phenotypic characterisation of a human iPSC-derived tri-culture using ioGlutamatergic Neurons, ioAstrocytes, and ioMicroglia
Using our fully optimised protocol, ioGlutamatergic Neurons (MAP2, red), ioMicroglia (IBA1, yellow) and ioAstrocytes (vimentin, cyan) were co-cultured to create a highly defined CNS model. High-resolution ICC analysis confirms the successful co-localisation and morphological health of three distinct cell types within a unified environment. By day 7, the protocol yields a highly consistent, integrated network suitable for complex cell modelling. DAPI (blue) highlights the total cell density and integrity of the culture. This protocol is compatible with derivative products of the three cell types, ensuring straightforward implementation across experimental workflows.

Efficient mRNA transfection into ioGlutamatergic Neurons
ioGlutamatergic Neurons are efficiently transfected and show sustained long-term expression of mRNA encoding GFP. ioGlutamatergic Neurons were imaged from day 1 post-thaw and throughout the experiment to assess transfection efficiency and evaluate potential cytotoxic effects of the transfection protocol. Day 1 images were captured prior to transfection on the same day.
Download the step-by-step protocol for lipid-based delivery of synthetic mRNA into ioGlutamatergic Neurons.

Lipid-based delivery of synthetic mRNA into ioGlutamatergic Neurons
ioGlutamatergic Neurons were transfected 24 hours post-thaw using Lipofectamine™ Stem Transfection Reagent. The transfection efficiency was evaluated by fluorescence imaging over 18 days after mRNA delivery, resulting in high transfection efficiency (close to 100%) and long-term sustained GFP expression.
Quantification of the GFP signal shows a decrease in GFP intensity over time, while the percentage of GFP-positive cells remains largely unchanged over time.
(A) The percentage of GFP-positive cells from two independent experiments.
(B) GFP intensity, quantified in successfully transfected cells from two independent experiments is quantified and normalised to day 2 (24 hours post-transfection).

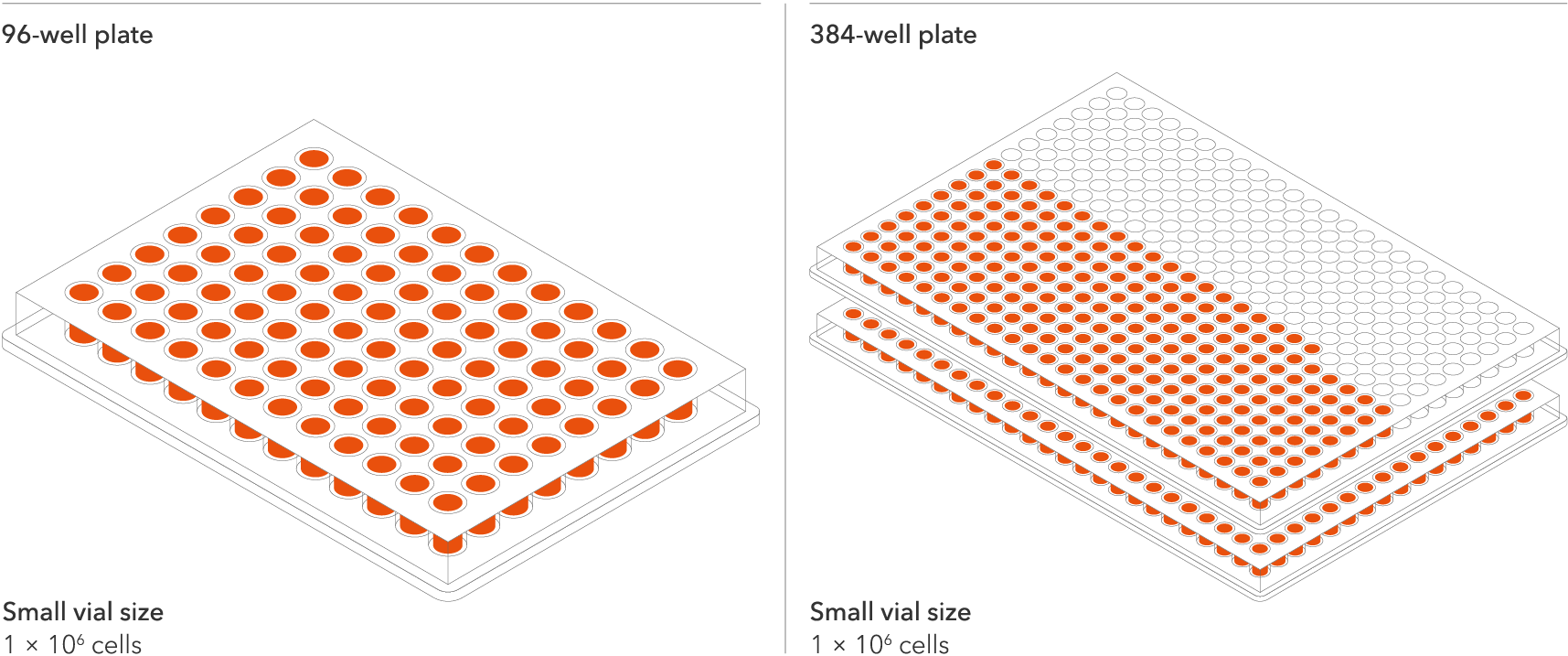
Industry leading seeding density
The recommended minimum seeding density is 30,000 cells/cm2, compared to up to 250,000 cells/cm2 for other similar commercially available products. One small vial can plate a minimum of 0.7 x 24-well plate, 1 x 96-well plate, or 1.5 x 384-well plates. This means every vial goes further, enabling more experimental conditions and more repeats, resulting in more confidence in the data.
Vial limit exceeded
A maximum number of 20 vials applies. If you would like to order more than 20 vials, please contact us at orders@bit.bio.





Hoescht(blue)TUBB3(blue)_day4.jpg?width=604&name=bit.bio_ioGlutamatergic%20Neurons_60xMAP2(red)Hoescht(blue)TUBB3(blue)_day4.jpg)

.png?width=1860&height=1260&name=bit.bio_3x2_ioGlutamatergic%20Neurons_MAP2_Hoescht_x20_hi.res%20(1).png)









