

.png?width=3430&height=2442&name=ioSkeletal%20Myocytes%20DMD%20Ex%2044%20DMD%20ICC%20combined%20(1).png)








.png?width=3430&height=2442&name=ioSkeletal%20Myocytes%20DMD%20Ex%2044%20DMD%20ICC%20combined%20(1).png)






cat no | io1018
ioSkeletal Myocytes DMD Exon 44 Deletion
Human iPSC-derived Duchenne muscular dystrophy model
-
Cryopreserved human iPSC-derived cells powered by opti-ox that are ready for experiments in days
-
Study Duchenne muscular dystrophy in a human in vitro model engineered with a DMD exon 44 deletion
-
Deletion results in weaker contraction and fatigue; dystrophin restored by ASO-mediated exon skipping

Human iPSC-derived Duchenne muscular dystrophy model

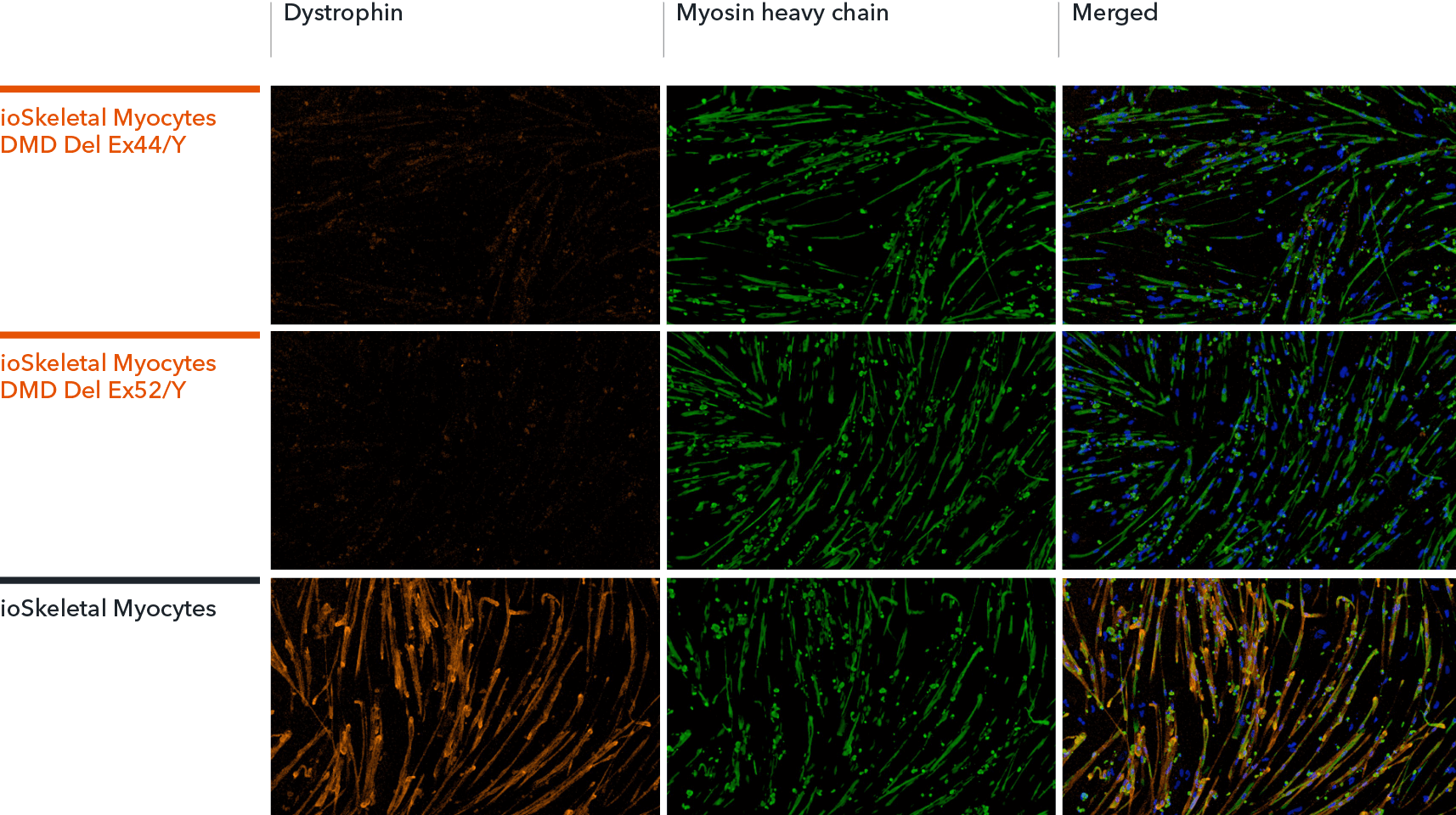
ioSkeletal Myocytes DMD Exon Deletion disease model cells show absence of Dystrophin protein by immunocytochemistry, demonstrating a Duchenne muscular dystrophy phenotype
ioSkeletal Myocytes DMD Exon 44 Deletion and DMD Exon 52 Deletion disease model cells, and ioSkeletal Myocytes wild type genetically matched control, were cultured in 96-well plates at a density of 32,000 cells per well, according to the user manual. Immunocytochemistry staining for Dystrophin and Myosin Heavy Chain (MHC) was carried out at day 10 post-revival. The data show robust expression of MHC, but no expression of Dystrophin in the ioSkeletal Myocytes DMD Del Ex44/Y or DMD Del Ex52/Y disease model cells compared to the wild-type control, demonstrating a Duchenne muscular dystrophy phenotype. Dystrophin was restored by ASO-mediated exon skipping. Data courtesy of Charles River Laboratories.
.png?width=3430&height=2442&name=ioSkeletal%20Myocytes%20DMD%20Ex%2044%20DMD%20ICC%20combined%20(1).png)
ioSkeletal Myocytes DMD Exon 44 Deletion disease model cells express skeletal muscle cell specific markers and lack expression of Dystrophin, demonstrating a Duchenne muscular dystrophy phenotype
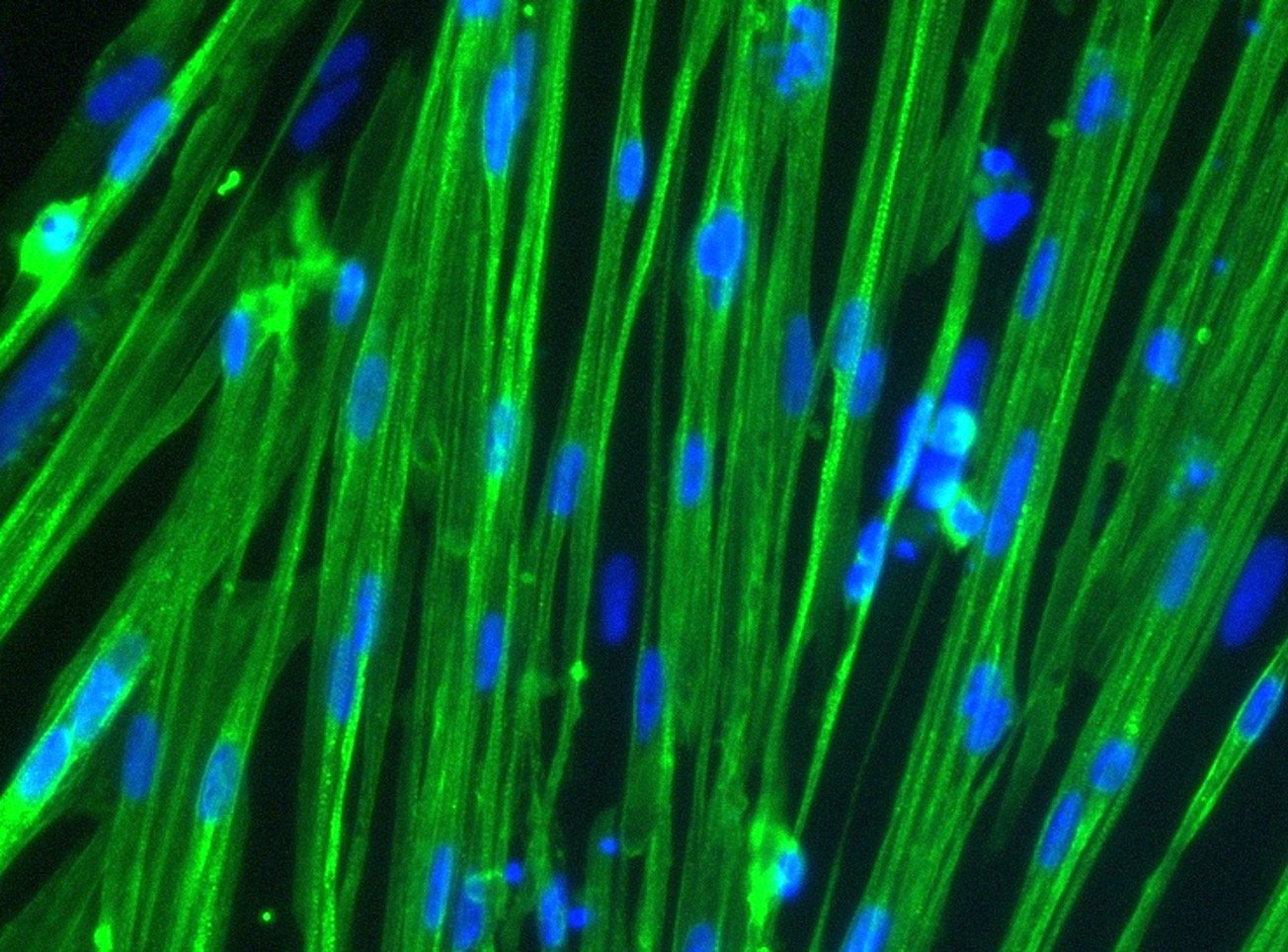
Immunocytochemistry staining at day 10 post revival demonstrates robust expression of Desmin, a component of the contractile apparatus, and no expression of Dystrophin in the ioSkeletal Myocytes DMD Del Ex44/Y disease model cells, whereas ioSkeletal Myocytes, the wild type isogenic control, express both markers (upper panel). Robust expression of Myosin Heavy Chain (MHC) and the muscle transcription factor Myogenin is observed in both ioSkeletal Myocytes DMD Del Ex44/Y and ioSkeletal Myocytes (lower panel). Anti-dystrophin antibody clone 2C6 (MANDYS106).

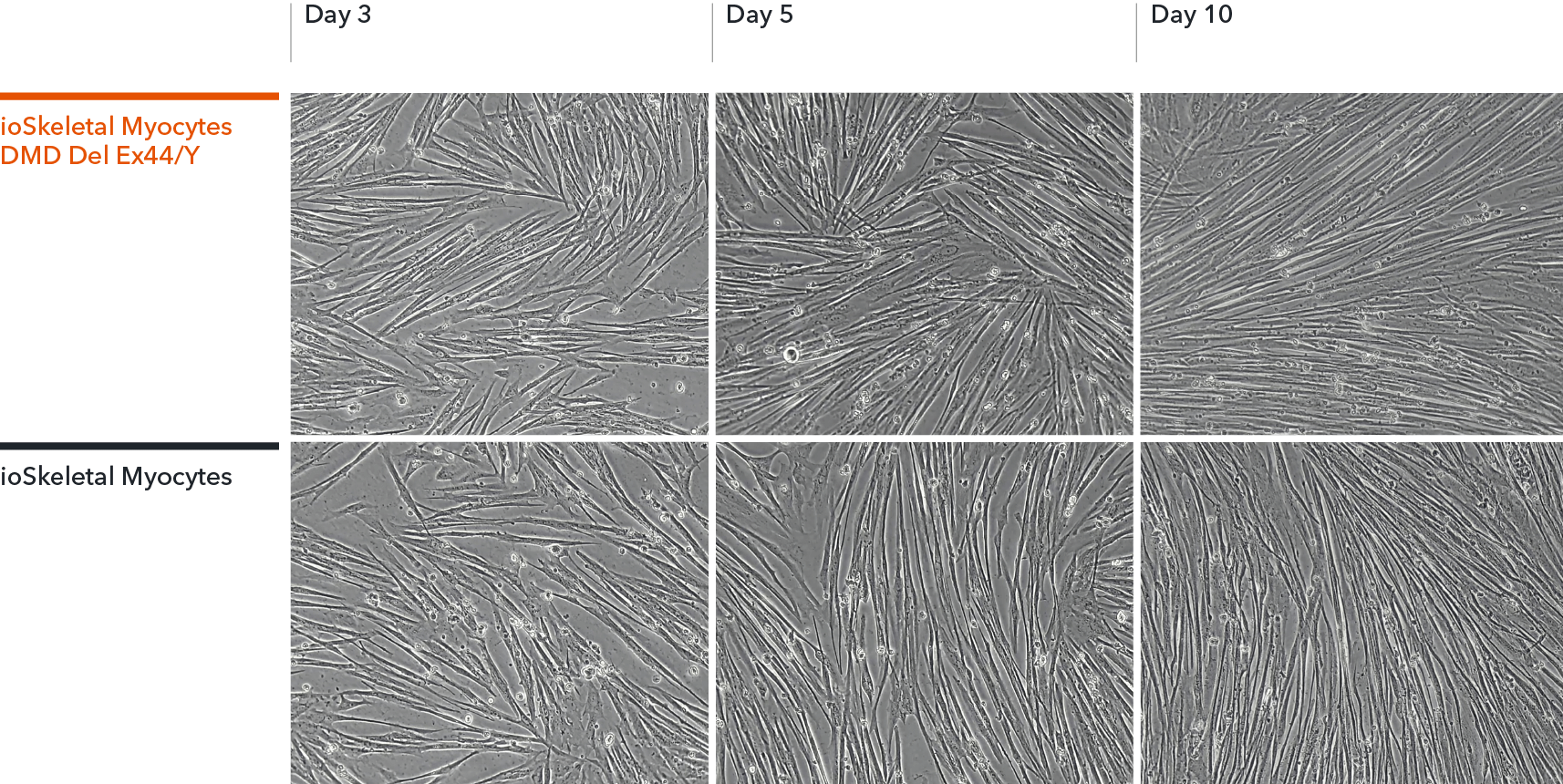
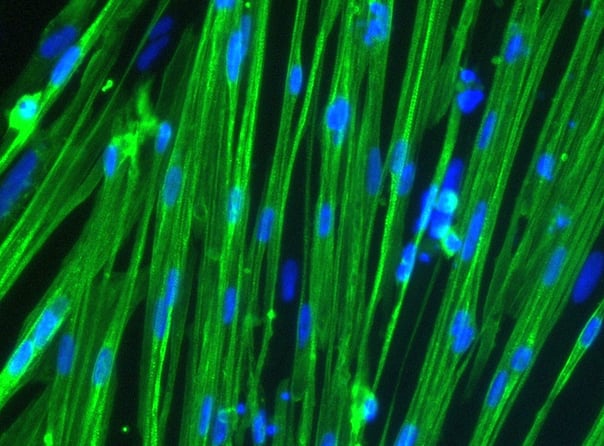
ioSkeletal Myocytes DMD Exon 44 Deletion disease model cells demonstrate classical skeletal myocytes morphology
ioSkeletal Myocytes DMD Exon 44 Deletion form elongated, multinucleated myocytes over 10 days, comparable to the wild-type ioSkeletal Myocytes isogenic control. Day 3 to 10 post-revival; 100X magnification.

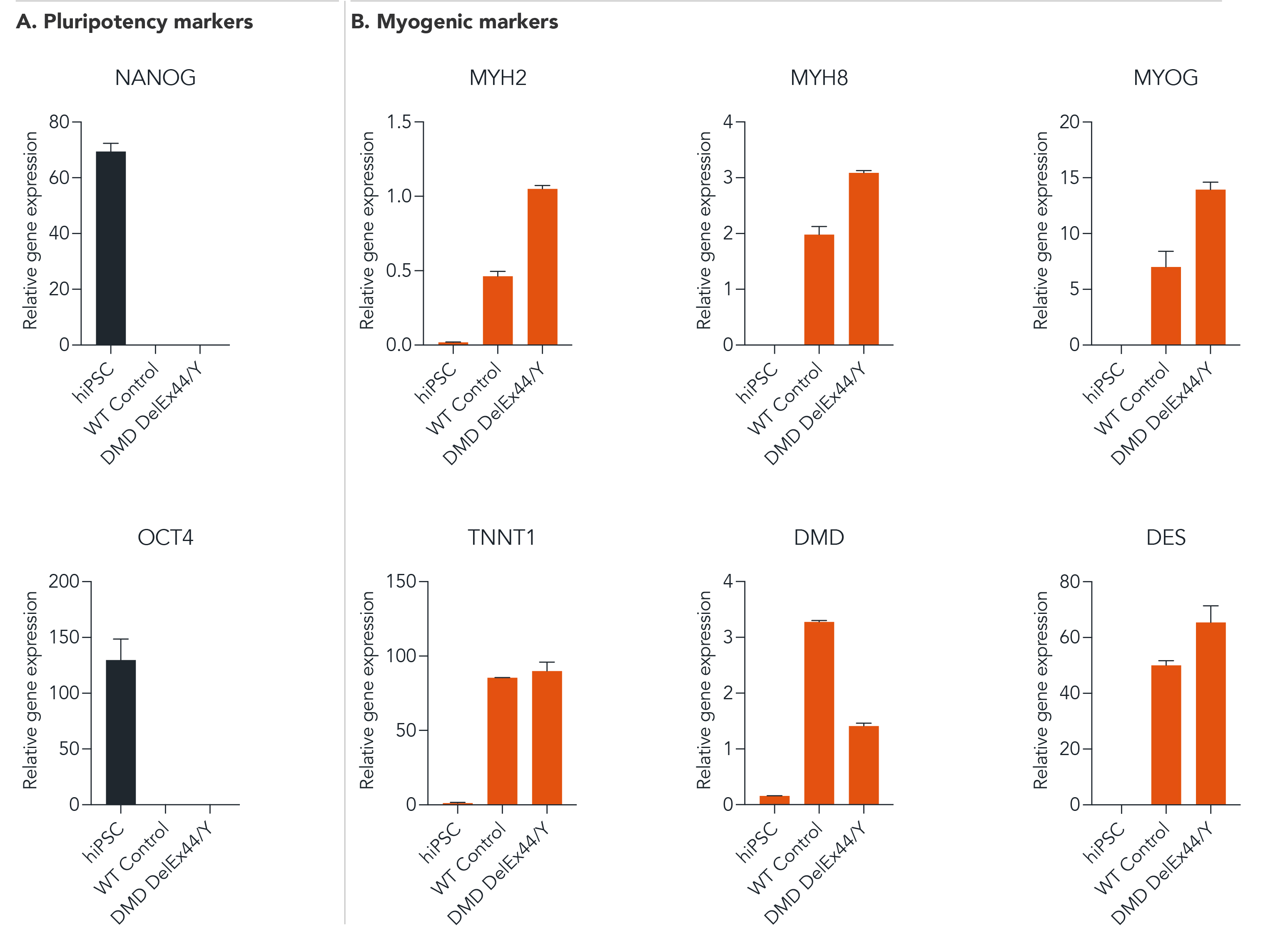
ioSkeletal Myocytes DMD Exon 44 Deletion disease model cells demonstrate gene expression of key myogenic markers following deterministic programming
Following reprogramming, ioSkeletal Myocytes DMD Exon 44 Deletion (DMD DelEx44/Y) and wild type ioSkeletal Myocytes (WT Control) downregulate expression of pluripotency genes (A), while demonstrating expected expression of key myogenic markers (B). Gene expression levels assessed by RT-qPCR. Data expressed relative to the parental human iPSC (hiPSC), normalised to HMBS. Data represents day 10 post-revival samples.
View the step-by-step RNA extraction and RT-qPCR protocol used to generate this data

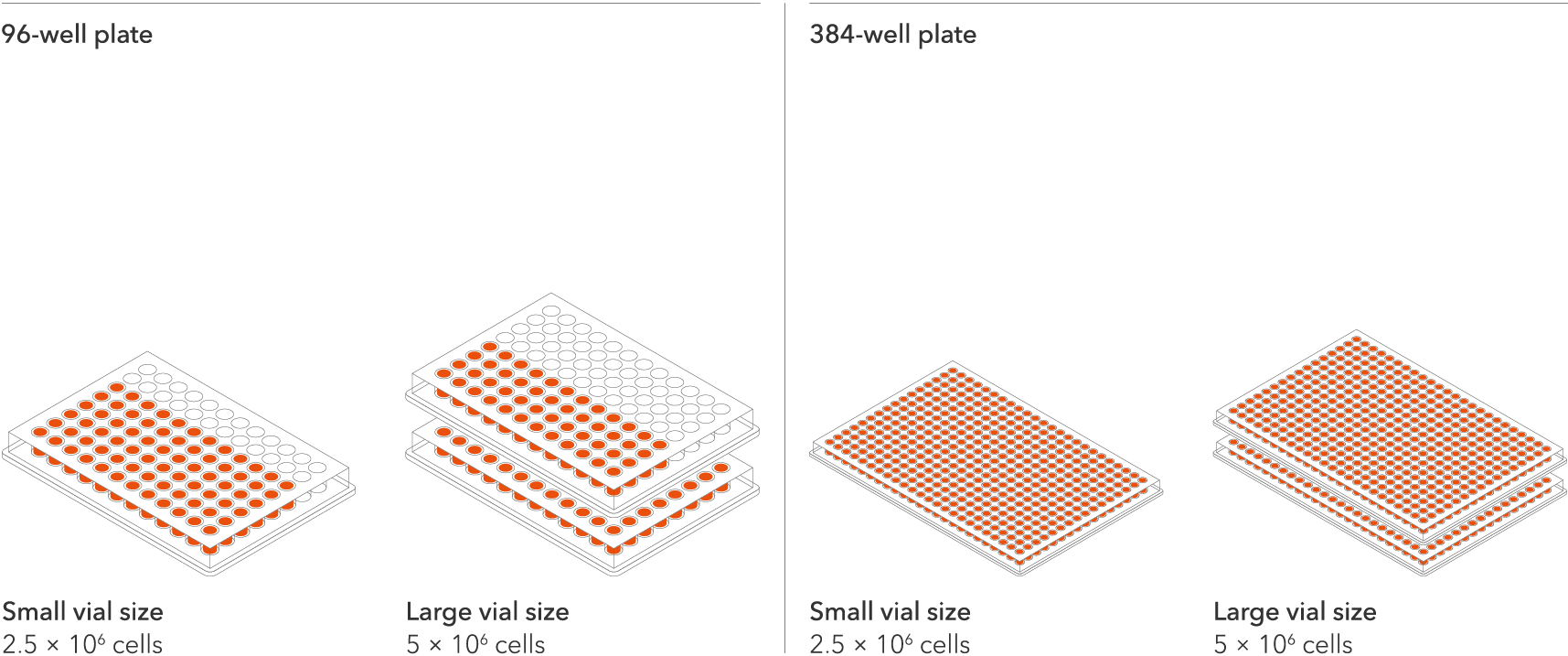
Industry leading seeding density
ioSkeletal Myocytes DMD Del Ex44/Y are compatible with plates ranging from 6 to 384 wells and are available in two vial sizes, tailored to suit your experimental needs with minimal waste.
The recommended seeding density is 100,000 cells/cm2.
One small vial can plate a minimum of 0.5 x 24-well plate, 0.75 x 96-well plate, or 1 x 384-well plate. One large vial can plate a minimum of 1 x 24-well plate, 1.5 x 96-well plates, or 2 x 384-well plates.

ASO-mediated Dystrophin restoration demonstrated in ioSkeletal Myocytes DMD Exon 44 Deletion disease model cells
ioSkeletal Myocytes (wild type control) and ioSkeletal Myocytes DMD Exon 44 Deletion (DMD Del Ex44) were cultured according to the user manual in 96-well plates at a density of 32,000 cells per well. On day 4 post-revival the cells were treated by gymnosis with exon 45 skipping antisense oligonucleotide (ASO-1), concentration range 1-50 µM for immunocytochemistry and high content image (HCI) analysis, and 0.01-50 µM for ddPCR. Cells were cultured to day 7 then analysed.
(A) Dystrophin mRNA restoration: RNA was extracted and cDNA synthesised. PCR primers and a fluorescent (FAM) labelled probe were designed to amplify the region coding exons 43-45 (non-skip, NSKP) or exons 43-46 (skip, SKP). PCR was carried out to quantify the non-skip and skip transcripts. The graph shows a concentration-dependent increase in the amount of SKP transcript (blue) and a decrease in the NSKP transcript (yellow), indicating that ASO-1 treatment has been successful in creating an in frame mRNA transcript for dystrophin.
(B) Dystrophin protein restoration: cells were stained for dystrophin, myosin heavy chain (not shown) and DAPI for cell nuclei quantification (not shown) and measured by high content analysis of images captured on Yokogawa Cell Voyager 8000 using a proprietary algorithm. The graph shows dystrophin restoration levels calculated as % of dystrophin/MHC area compared to wild type. ASO-1 treatment increased dystrophin protein expression in a concentration-dependent manner.
Each condition was tested in technical duplicate for ddPCR and technical triplicate for HCI analysis, and in biological duplicate (N=2). Data courtesy of Charles River Laboratories.

DMD Exon Deletion 3D muscle microtissues show weaker contraction and fatigue
Wild-type (WT) and DMD exon deletion disease model ioSkeletal Myocytes were cultured in 3D on the MUSbit microchip (Bi/ond), which includes pillars designed for anchoring muscle cell bundles. Muscle microtissues developed over 14 days.
When compared to the genetically matched control (WT) at day 14, the DMD Exon 44 Deletion and Exon 52 Deletion models showed weaker contraction upon twitch and tetanus stimuli (A), and fatigue under sustained stimulation (B).
The 3D muscle microtissues provide a relevant model to study how exon deletions in the dystrophin gene affect muscle contraction.
Data courtesy of M. Han and M. Aarts, formerly at Bi/ond Solutions BV.

Co-culture of ioSkeletal Myocytes and ioMotor Neurons
High resolution confocal imaging of ioSkeletal Myocytes (io1002) and ioMotor Neurons (io1027) co-culture. Staining with alpha-bungarotoxin (yellow) highlights acetylcholine receptor expression on co-cultured ioSkeletal Myocytes. Desmin (cyan) and microtubule-associated protein 2 (red) define ioSkeletal Myocytes and ioMotor Neurons respectively. Co-culture imaged at day 30, 40X magnification.
Download the step-by-step protocol for culturing ioMotor Neurons and ioSkeletal Myocytes.
Vial limit exceeded
A maximum number of 20 vials applies. If you would like to order more than 20 vials, please contact us at orders@bit.bio.

.png?width=3430&height=2442&name=ioSkeletal%20Myocytes%20DMD%20Ex%2044%20DMD%20ICC%20combined%20(1).png)


.png?width=604&name=ISSCR24-DMD-Exon44-3D-muscle-bundle%20(1).png)





/ioSkeletal-Myocytes-DMD-Exon-52-Deletion-hero.webp?width=1276&height=1190&name=ioSkeletal-Myocytes-DMD-Exon-52-Deletion-hero.webp)


