cat no | io1094
CRISPRko-Ready ioMicroglia Male
Male human iPSC donor-derived microglia expressing Cas9 for rapid gene knockout generation
-
Cryopreserved human iPSC-derived cells powered by opti-ox, that are ready for experiments in days
-
Microglia constitutively expressing Cas9 optimised for single/combinatorial gene knockouts and CRISPR screens
-
Consistently perform key phagocytic and cytokine secretion functions, and are co-culture compatible

Male human iPSC donor-derived microglia expressing Cas9 for rapid gene knockout generation

Flow cytometry analysis demonstrates high knockout efficiency of beta-2 microglobulin (B2M) by both lentiviral transduction and lipid-based transfection
Flow cytometry analysis of B2M protein expression in CRISPRko-Ready ioMicroglia, after delivery of gRNA targeting B2M. gRNAs were introduced into the cells at day 10 post-thaw using two methods: lentiviral transduction or synthetic gRNA delivery with Lipofectamine RNAiMAX transfection reagent. After 5 days of culture following guide delivery, B2M gene knockout efficiency was assessed by flow cytometry analysis. (A) Lentiviral transduction with gRNA targeting B2M: 20% of cells received a B2M gRNA, as measured by GFP expression. A high knockout efficiency of 86% was achieved in these GFP+ cells (B). (C) Lipid-based transfection with gRNA targeting B2M: a high knockout efficiency of 82% was achieved.

A pooled single cell CRISPR knockout screen uncovers modulators of microglia activation
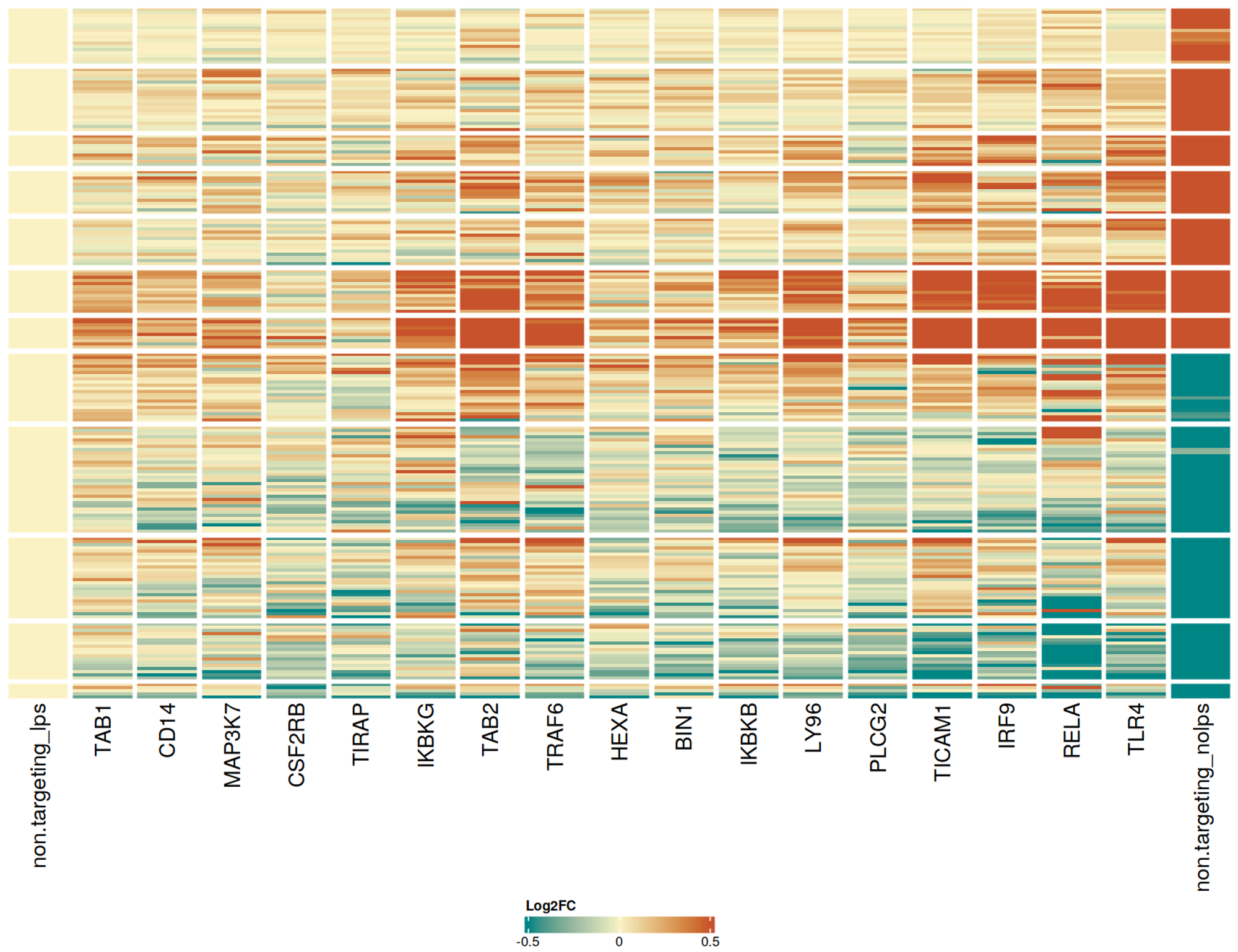
To conduct a pooled single cell CRISPR knockout screen (scCRISPR screen) with a targeted sequencing readout, we first identified a transcriptomic activation signature of 258 differentially expressed genes by comparing CRISPRko-Ready ioMicroglia treated with and without LPS at day 10. Separately, we selected 110 candidate genes for the pooled scCRISPR screen based on their known roles in neurodegeneration and neuroinflammation. Guide RNAs were delivered via lentiviral transduction on day 10, aiming for a single integration per cell. The cells were cultured and then treated with +/- LPS for 24 hours before single cell processing on day 15. Cosine similarity analysis compared knockouts in LPS-treated CRISPRko-Ready ioMicroglia to both resting and activated states. The analysis identified 17 gene knockouts that altered responses to LPS stimulation. The heatmap shows Log2FC profiles for gene knockouts that had a cosine similarity above 0.3 (arbitrarily chosen threshold) compared to cells with non-targeting guides in the unstimulated condition. Knockouts are sorted based on their cosine similarity to the non-LPS condition. CD14, MAP3K7, TIRAP, IKBKG, TRAF6, IKBKB, LY96, TICAM1, RELA, and TLR4 are genes known to be involved in LPS activation mediated via the TLR4 signalling pathway.

Phagocytosis of E. coli particles by CRISPRko-Ready ioMicroglia
(A) Phagocytosis assay using pHrodo™ E. coli BioParticles™ at day 10 post-thaw demonstrates efficient uptake of bacteria particles by CRISPRko-Ready ioMicroglia in comparison to ioMicroglia Male (io1021) after 4 hours of treatment. (B) An increase of fluorescence intensity of E.coli particles upon pH change in the phagosome, can be readily detected by fluorescence microscopy. A steep increase of fluorescence signal intensity was measured in the presence of E.coli particles alone, but not in combination with Cytochalasin D (CytoD), an inhibitor of actin polymerization.

CRISPRko-Ready ioMicroglia secrete pro-inflammatory cytokines upon activation
CRISPRko-Ready ioMicroglia were stimulated at day 10 post-thaw with LPS 100 ng/ml and IFNɣ 20 ng/ml for 24 hours. Supernatants were harvested and analysed by quantitative ELISA. These cells secrete the pro-inflammatory cytokines, TNF⍺ and IL-6 upon activation in comparison to ioMicroglia Male (io1021).

CRISPRko-Ready ioMicroglia show ramified morphology by day 10
CRISPRko-Ready ioMicroglia mature rapidly and key ramified morphology can be identified by day 4 and continues through to day 10, similarly to ioMicroglia Male (io1021). Day 1 to 10 post-thawing; 100x magnification.

CRISPRko-Ready ioMicroglia express key microglia markers
Immunofluorescent staining on day 10 post-revival demonstrates similar homogenous expression of microglia marker P2RY12 and ramified morphology in CRISPRko-Ready ioMicroglia compared to ioMicroglia Male (io1021). 100X magnification.

CRISPRko-Ready ioMicroglia express key microglia markers
Immunofluorescent staining on day 10 post-revival demonstrates similar homogenous expression of microglia marker IBA1 and ramified morphology in CRISPRko-Ready ioMicroglia compared to ioMicroglia Male (io1021). 100X magnification.
Vial limit exceeded
A maximum number of 20 vials applies. If you would like to order more than 20 vials, please contact us at orders@bit.bio.











 Emmanouil Metzakopian
Emmanouil Metzakopian