Discover CRISPR-Ready
ioGlutamatergic Neurons™
Generate gene knockouts in hiPSC-derived neurons in days
CRISPR-Ready ioGlutamatergic Neurons are human iPSC-derived cells powered by opti-ox™, designed for scientists looking to generate gene knockouts in a physiologically relevant human neuronal background. Avoid lengthy stable Cas9 iPSC line engineering and characterisation, and rapidly generate gene knockouts in days.
The cells arrive ready to use. Using bit.bio’s simple cell culturing protocol with optimised guide RNA (gRNA) delivery, users can achieve high gene knockout efficiencies, and obtain functional experimental readouts from 4 days post-thaw. By following our simple optimised cell culture protocols, achieve robust experimental readouts without prior expertise in gRNA delivery optimisation in human neurons.
Benchtop benefits
Ready to use
Defined and characterised human neurons constitutively expressing Cas9, ready for knockout experiments from day 1.
Quick and easy
Generate readouts within days using a simple protocol for cell maturation and guide RNA delivery.
High knockout efficiency
Optimised protocols for lipid or lentivirus based guide RNA delivery ensure maximal knockout efficiency.
Go from seeding to knockout to readout in days
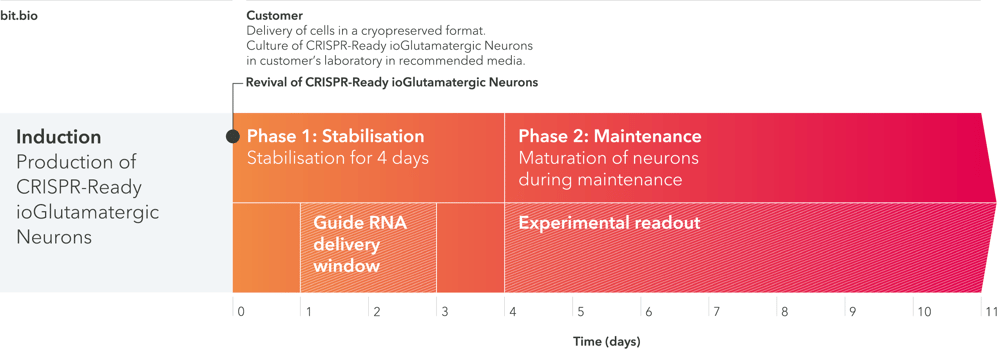

Start measuring the impact of knocking out your gene of interest in 4 simple steps
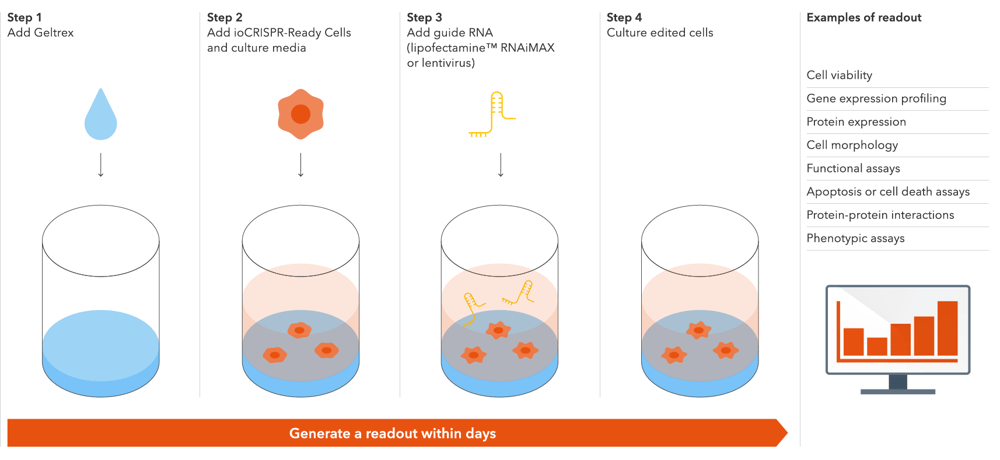
Our optimised protocol leads to high knockout efficiency with lentiviral transduction and lipid-based transfection
Using our optimised protocols for gRNA delivery, similar knockout efficiencies of SOX11 were achieved for both lentiviral transduction and lipid-based transfection.
SOX11 indel formation was measured by amplicon sequencing of CRISPR-Ready ioGlutamatergic Neurons transfected or transduced with a gRNA targeting SOX11. gRNAs were delivered at either day 1 or day 3 after thawing by either lentiviral transduction or as a synthetic gRNA using Lipofectomine™ RNAiMAX transfection reagent. DNA was harvested from cells after three days of culture post-transduction and transfection.
![]()
Get neurons at scale for CRISPR screens
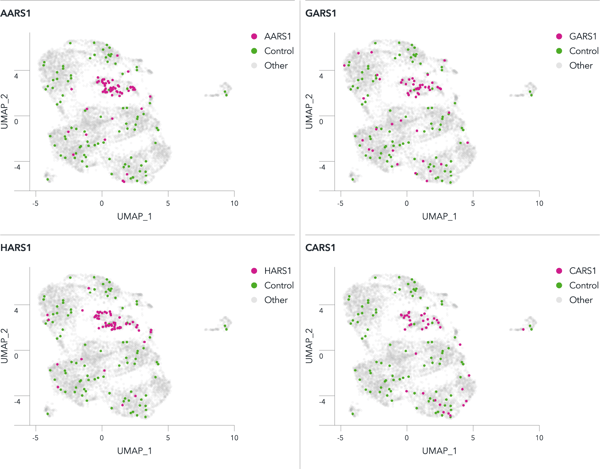
Use CRISPR-Ready ioGlutamatergic Neurons for knockout screens in a physiologically relevant background.
A pooled knockout screen of neurodegenerative disease-relevant genes in CRISPR-Ready ioGlutamatergic Neurons shows clustering of aaRS genes in UMAPs
We selected 100 genes involved in neurodegenerative diseases for a pooled CRISPR screen. Lentiviral transduction of the gRNAs was carried out at day 3, and single cell gene expression analysis was performed at day 12. Clustering of aminoacyl-tRNA synthetase (aaRSs) knockouts including AARS1, HARS1, CARS1, and GARS1 was observed. In contrast, cells transduced with non-targeting control sgRNAs were evenly distributed among clusters. Pathway analysis showed gRNAs targeting aaRSs activated the unfolded protein response, a pathway of significance for neurodegenerative diseases.
Perform large-scale screens in cells that closely resemble their in vivo counterparts
As ioCRISPR-Ready Cells are powered by opti-ox precision cell reprogramming, they have consistency and scalability built-in. With these cells, users can significantly cut experimental timelines by no longer needing to spend months engineering and characterising their own Cas9 stable iPSC lines or optimising differentiation protocols.
“To do a genome-level CRISPR screen, with all the necessary replicates, requires billions of cells. Reaching that scale with iPSCs has been a significant challenge, so many people turn to immortalised cell lines. But these cells are quite different from neurons in the human body. The development of ioCRISPR-Ready Cells is a huge step forward because it allows us to perform large-scale CRISPR screens on cells that closely resemble their in vivo counterparts — it’s a more physiologically relevant way of doing things.”
Emmanouil Metzakopian, PhD
VP of Research and development, bit.bio
Looking to test a vial of CRISPR-Ready ioGlutamatergic Neurons?
We are seeking scientists looking for early access to CRISPR-Ready ioGlutamatergic Neurons. Fill out the form to register your interest now, and our team will be in touch with further details about our Early Access Program.
Want to learn more?
